In the last part (here) we setup a simple R install so we could look at analysing and plotting perfmon data in R. In this post we’ll look about creating a very simple plot from a perfmon CSV. In later posts I’ll show some examples of how to clean the data up, to pull it from a SQL Server repository, combine datasets for analysis and some of the other interesting things R lets you do.
So lets start off with some perfmon data. Here’s a CSV (R-perfmon) that contains the following counters:
- Physical Disk C:\ Average Disk Queue Length
- Physical Disk C:\ Average Disk Read Queue Length
- Physical Disk C:\ Average Disk Write Queue Length
- % Processor Time
- Processes
- Processor Queue Length
Perfmon was set to capture data every 15 seconds.
Save this to somewhere. For the purposes of the scripts I’m using I’ll assume you’ve put it in the folder c:\R-perfmon.
Fire up your R environment of choice, I’ll be using R Studio. So opening a new instance, and I’m granted by a clean workspace:
On the left hand side I’ve the R console where I’ll be entering the commands and on the right various panes that left me explore the data and graphs I’ve created.
As mentioned before R is a command line language, it’s also cAse Sensitive. So if you get any strange errors while running through this example it’s probably worth checking exactly what you’ve typed. If you do make a mistake you can use the cursor keys to scroll back through commands, and then edit the mistake.
So the first thing we need to do is to install some packages, Packages are a means of extending R’s capabilities. The 2 we’re going to install are ggplot2 which is a graphing library and reshape2 which is a library that allows us to reshape the data (basically a Pivot in SQL Server terms). We do this with the following command:
install.packages(c("ggplot2","reshape2"))
You may be asked to pick a CRAN mirror, select the one closest to you and it’ll be fine. Assuming everything goes fine you should be informed that the packages have been installed, so they’ll now be available the next time you use R. To load them into your current session, you use the commands:
library("ggplot2")
library("reshape2")
So that’s all the basic housekeeping out of the way, now lets load in some Perfmon data. R handles data as vectors, or dataframes. As we have multiple rows and columns of data we’ll be loading it into a dataframe.
data <-read.table("C:\\R-perfmon\\R-perfmon.csv",sep=",",header=TRUE)
IF everything’s worked, you’ll see no response. What we’ve done is to tell R to read the data from our file, telling it we’re using , as the seperator and that the first row contains the column headers. R using the ‘<-‘ as an assignment operator.
To prove that we’ve loaded up some data we can ask R to provide a summary:
summary(data)
X.PDH.CSV.4.0...GMT.Daylight.Time...60.
04/15/2013 00:00:19.279: 1
04/15/2013 00:00:34.279: 1
04/15/2013 00:00:49.275: 1
04/15/2013 00:01:04.284: 1
04/15/2013 00:01:19.279: 1
04/15/2013 00:01:34.275: 1
(Other) :5754
X..testdb1.PhysicalDisk.0.C...Avg..Disk.Queue.Length
Min. :0.000854
1st Qu.:0.008704
Median :0.015553
Mean :0.037395
3rd Qu.:0.027358
Max. :4.780562
X..testdb1.PhysicalDisk.0.C...Avg..Disk.Read.Queue.Length
Min. :0.000000
1st Qu.:0.000000
Median :0.000980
Mean :0.017626
3rd Qu.:0.003049
Max. :4.742742
X..testdb1.PhysicalDisk.0.C...Avg..Disk.Write.Queue.Length
Min. :0.0008539
1st Qu.:0.0076752
Median :0.0133689
Mean :0.0197690
3rd Qu.:0.0219051
Max. :2.7119064
X..testdb1.Processor._Total....Processor.Time X..testdb1.System.Processes
Min. : 0.567 Min. : 77.0
1st Qu.: 7.479 1st Qu.: 82.0
Median : 25.589 Median : 85.0
Mean : 25.517 Mean : 87.1
3rd Qu.: 38.420 3rd Qu.: 92.0
Max. :100.000 Max. :110.0
X..testdb1.System.Processor.Queue.Length
Min. : 0.0000
1st Qu.: 0.0000
Median : 0.0000
Mean : 0.6523
3rd Qu.: 0.0000
Max. :58.0000
And there we are, some nice raw data there. Some interesting statistical information given for free as well. Looking at it we can see that our maximum Disk queue lengths aren’t anything to worry about, and even though our Processor peaks at 100% utilisation, we can see that it spends 75% of the day at less the 39% utilisation. And we can see that our Average queue length is nothing to worry about.
But lets get on with the graphing. At the moment R doesn’t know that column 1 contains DateTime information, and the names of the columns are rather less than useful. To fix this we do:
cname<-c("Time","Avg Disk Queue Length","Avg Disk Read Queue Length","Avg Disk Write Queue Length","Total Processor Time%","System Processes","System Process Queue Length")
colnames(data)<-cname
data$Time<-as.POSIXct(data$Time, format='%m/%d/%Y %H:%M:%S')
mdata<-melt(data=data,id.vars="Time")
First we build up an R vector of the column names we’d rather use, “c” is the constructor to let R know that the data that follows is to interpreted as vector. Then we pass this vector as an input to the colnames function that renames our dataframe’s columns for us.
On line 3 we convert the Time column to a datetime format using the POSIXct function and passing in a formatting string.
Line 4, we melt our data. Basically we’re turning our data from this:
| Time | Variable A | Variable B | Variable C |
|---|---|---|---|
| 19/04/2013 14:55:15 | A1 | B2 | C9 |
| 19/04/2013 14:55:30 | A2 | B2 | C8 |
| 19/04/2013 14:55:45 | A3 | B2 | C7 |
to this:
| ID | Variable | Value |
|---|---|---|
| 19/04/2013 14:55:15 | Variable A | A1 |
| 19/04/2013 14:55:30 | Variable A | A2 |
| 19/04/2013 14:55:45 | Variable A | A3 |
| 19/04/2013 14:55:15 | Variable B | B2 |
| 19/04/2013 14:55:30 | Variable B | B2 |
| 19/04/2013 14:55:45 | Variable B | B2 |
| 19/04/2013 14:55:15 | Variable C | C9 |
| 19/04/2013 14:55:30 | Variable C | C8 |
| 19/04/2013 14:55:45 | Variable C | C7 |
This allows to very quickly plot all the variables and their values against time without having to specify each series
ggplot(data=mdata,aes(x=Time,y=value,colour=variable))+ geom_point(size=.2) + stat_smooth() + theme_bw()
This snippet introduces a new R technique. By putting + at the end of the line you let R know that the command is spilling over to another line. Which makes complex commands like this easier to read and edit. Breaking it down line by line:
- tells R that we want to use ggplot to draw a graph. The data parameter tells ggplot which dataframe we want to plot.
aeslets us pass in aesthetic information, in this case we tell it that we want Time along the x axis, the value of the variable on the y access, and to group/colour the values by variable - tells ggplot how large we’d like the data points on the graph.
- This tells we want ggplot to draw a best fit curve through the data points.
- Telling ggplot which theme we’d like it to use. This is a simple black and white theme.
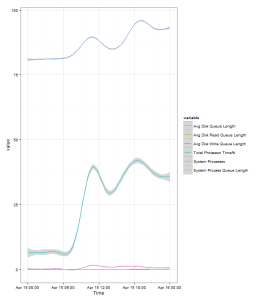
Run this and you’ll get:
The “banding” in the data points for the System processes count is due to the few discrete values that the variable takes.
The grey banding around the fitted line is a 95% confidence interval. The wider it is the greater variance of values there are at that point, but in this example it’s fairly tight to the plot so you can assume that it’s a good fit to the data.
We can see from the plot that the Server is ticking over first thing in the morning then we see an increases in load as people start getting into the office from 07:00 onwards. Appears to be a drop off over lunch as users head away from their clients, and then picks up through the afternoon. Load stays high through the rest of the day, and in this example it ends higher than it started as there’s an overnight import running that evening, though if you hadn’t know that this plot would have probably raised questions and you’d have investigate. Looking at the graph it appears that all the load indicators (Processor time% and System Processes) follow each other nicely, later in this series we’ll look at analysing which one actually leads the other
You can also plot the graph without all the data points for a cleaner look:
ggplot(data=mdata,aes(x=Time,y=value,colour=variable))+ + stat_smooth() + + theme_bw()
So putting all that together we have the following script which can just be cut and pasted across into R:
install.packages(c("ggplot2","reshape2"))
library("ggplot2")
library("reshape2")
data <-read.table("C:\\R-perfmon\\R-perfmon.csv",sep=",",header=TRUE)
cname<-c("Time","Avg Disk Queue Length","Avg Disk Read Queue Length","Avg Disk Write Queue Length","Total Processor Time%","System Processes","System Process Queue Length")
colnames(data)<-cname
data$Time<-as.POSIXct(data$Time, format='%m/%d/%Y %H:%M:%S')
mdata<-melt(data=data,id.vars="Time")
ggplot(data=mdata,aes(x=Time,y=value,colour=variable))+
geom_point(size=.2) +
stat_smooth() +
theme_bw()



Stephen
Wow. Super cool! Keep up the great work!!
Rajesh
Does not work for me. Below is the error message
Error in seq.int(0, to0 – from, by) : ‘to’ cannot be NA, NaN or infinite
In addition: Warning message:
Removed 34560 rows containing non-finite values (stat_smooth).
Stuart Moore
Check your perfmon data. That error message sounds like you’ve got some nulls or non-numeric (NaN == Not a Number) data in the frame and that’s causing smooth to throw an error.